library(dplyr)
library(tidyr)
library(ggplot2)
library(metafor)
library(gsheet)
library(tidytext)
library(cowplot)Packages
gs_url <- "https://docs.google.com/spreadsheets/d/1i7f61MP5efJclMLpQWgh9k3iTyX7Xgvj/edit?gid=1609934999#gid=1609934999"diseases <- c("poli","comum","branca","cercos","turc","diplo","antrac","bipol")
thr_present <- 1.6
price <- 200
cost <- 80 dat <- gsheet2tbl(gs_url) %>%
mutate(
application = tolower(trimws(application)),
season = factor(tolower(trimws(season)), levels = c("first","second")),
location = trimws(location),
trial = paste(year, location, season, sep = "_")
)
stopifnot(all(c("check","treated") %in% dat$application))
dat$poli[is.na(dat$poli)] <- 0
dat$comum[is.na(dat$comum)] <- 0
dat$branca[is.na(dat$branca)] <- 0
dat$cercos[is.na(dat$cercos)] <- 0
dat$turc[is.na(dat$turc)] <- 0
dat$diplo[is.na(dat$diplo)] <- 0
dat$antrac[is.na(dat$antrac)] <- 0
dat$bipol[is.na(dat$bipol)] <- 0hyb_app <- dat %>%
group_by(trial, year, location, season, hybrid, application) %>%
summarise(
yield = mean(yield, na.rm = TRUE),
across(all_of(diseases), mean, na.rm = TRUE),
.groups = "drop"
)
hyb_apphyb_diff <- hyb_app %>%
pivot_wider(
names_from = application,
values_from = c(yield, all_of(diseases))
) %>%
filter(!is.na(yield_check) & !is.na(yield_treated)) %>%
mutate(d_yld = yield_treated - yield_check)es <- hyb_diff %>%
rowwise() %>%
mutate(sev_sum_check = sum(c_across(all_of(paste0(diseases, "_check"))), na.rm = TRUE)) %>%
ungroup() %>%
group_by(trial, year, location, season) %>%
summarise(
yi = mean(d_yld, na.rm = TRUE),
vi = var(d_yld, na.rm = TRUE) / sum(!is.na(d_yld)),
m_hybrids = sum(!is.na(d_yld)),
sev_sum = mean(sev_sum_check, na.rm = TRUE),
.groups = "drop"
) %>%
mutate(
pressure = factor(
ifelse(sev_sum > thr_present, "Disease present (>2%)", "Disease absent (≤2%)"),
levels = c("Disease absent (≤2%)", "Disease present (>2%)")
)
) %>%
filter(is.finite(vi), vi > 0, m_hybrids >= 2)# %>%
#filter(!trial == "2019_Pato Branco_first")es %>%
filter(sev_sum > 0.1) %>%
summarise(
mean = mean(sev_sum),
min = min(sev_sum),
max = max(sev_sum),
median = median(sev_sum)
)m0 <- rma(yi, vi, data = es, method = "REML")
m0
Random-Effects Model (k = 19; tau^2 estimator: REML)
tau^2 (estimated amount of total heterogeneity): 0.3904 (SE = 0.1385)
tau (square root of estimated tau^2 value): 0.6249
I^2 (total heterogeneity / total variability): 95.53%
H^2 (total variability / sampling variability): 22.37
Test for Heterogeneity:
Q(df = 18) = 264.1120, p-val < .0001
Model Results:
estimate se zval pval ci.lb ci.ub
0.4456 0.1480 3.0117 0.0026 0.1556 0.7357 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1mP <- rma(yi, vi, mods = ~ pressure, data = es, method = "REML")
mP
Mixed-Effects Model (k = 19; tau^2 estimator: REML)
tau^2 (estimated amount of residual heterogeneity): 0.2623 (SE = 0.0985)
tau (square root of estimated tau^2 value): 0.5122
I^2 (residual heterogeneity / unaccounted variability): 93.36%
H^2 (unaccounted variability / sampling variability): 15.06
R^2 (amount of heterogeneity accounted for): 32.81%
Test for Residual Heterogeneity:
QE(df = 17) = 191.4251, p-val < .0001
Test of Moderators (coefficient 2):
QM(df = 1) = 9.0906, p-val = 0.0026
Model Results:
estimate se zval pval ci.lb
intrcpt 0.0926 0.1692 0.5469 0.5844 -0.2392
pressureDisease present (>2%) 0.7430 0.2464 3.0151 0.0026 0.2600
ci.ub
intrcpt 0.4243
pressureDisease present (>2%) 1.2260 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1mP2 <- rma(yi, vi, mods = ~ season, data = es, method = "REML")
mP2
Mixed-Effects Model (k = 19; tau^2 estimator: REML)
tau^2 (estimated amount of residual heterogeneity): 0.4033 (SE = 0.1470)
tau (square root of estimated tau^2 value): 0.6351
I^2 (residual heterogeneity / unaccounted variability): 95.62%
H^2 (unaccounted variability / sampling variability): 22.85
R^2 (amount of heterogeneity accounted for): 0.00%
Test for Residual Heterogeneity:
QE(df = 17) = 259.2878, p-val < .0001
Test of Moderators (coefficient 2):
QM(df = 1) = 0.5011, p-val = 0.4790
Model Results:
estimate se zval pval ci.lb ci.ub
intrcpt 0.3316 0.2205 1.5040 0.1326 -0.1005 0.7637
seasonsecond 0.2133 0.3013 0.7079 0.4790 -0.3772 0.8037
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1es$pressure <- factor(es$pressure,
levels = c("Disease absent (≤2%)", "Disease present (>2%)"))
newP <- data.frame(pressure = levels(es$pressure))
X_P <- model.matrix(~ pressure, data = newP)[, -1, drop = FALSE] # one column
pred_P <- predict(mP, newmods = X_P)
pred_tab <- data.frame(
pressure = newP$pressure,
gain_hat = pred_P$pred,
se = pred_P$se,
ci_lb = pred_P$ci.lb,
ci_ub = pred_P$ci.ub
)
pred_tabes$season <- factor(es$season,
levels = c("first", "second"))
newP <- data.frame(season = levels(es$season))
X_P <- model.matrix(~ season, data = newP)[, -1, drop = FALSE] # one column
pred_P <- predict(mP2, newmods = X_P)
pred_tab2 <- data.frame(
season = newP$season,
gain_hat = pred_P$pred,
se = pred_P$se,
ci_lb = pred_P$ci.lb,
ci_ub = pred_P$ci.ub
)
pred_tab2hyb_lnr <- hyb_diff %>%
filter(yield_check > 0, yield_treated > 0) %>%
mutate(lnRR = log(yield_treated / yield_check))
es_lnr <- hyb_lnr %>%
group_by(trial, year, location, season) %>%
summarise(
yi = mean(lnRR, na.rm = TRUE),
vi = var(lnRR, na.rm = TRUE) / sum(!is.na(lnRR)),
.groups = "drop"
) %>%
filter(is.finite(vi), vi > 0) %>%
left_join(es %>% select(trial, pressure), by = "trial")
es_lnrmP_lnr <- rma(yi, vi, mods = ~ pressure, data = es_lnr, method = "REML")
mP_lnr
Mixed-Effects Model (k = 19; tau^2 estimator: REML)
tau^2 (estimated amount of residual heterogeneity): 0.0042 (SE = 0.0016)
tau (square root of estimated tau^2 value): 0.0645
I^2 (residual heterogeneity / unaccounted variability): 93.11%
H^2 (unaccounted variability / sampling variability): 14.52
R^2 (amount of heterogeneity accounted for): 35.21%
Test for Residual Heterogeneity:
QE(df = 17) = 163.7050, p-val < .0001
Test of Moderators (coefficient 2):
QM(df = 1) = 9.5946, p-val = 0.0020
Model Results:
estimate se zval pval ci.lb
intrcpt 0.0122 0.0214 0.5692 0.5692 -0.0298
pressureDisease present (>2%) 0.0966 0.0312 3.0975 0.0020 0.0355
ci.ub
intrcpt 0.0542
pressureDisease present (>2%) 0.1577 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1# HIGH
s <- summary(mP_lnr)
# Extract the moderator row (difference)
b1_est <- s$b[2] # lnRR difference: present - absent
b1_lb <- s$ci.lb[2] # lower CI
b1_ub <- s$ci.ub[2] # upper CI
# Convert lnRR difference to ratio
ratio_diff <- exp(b1_est)
ratio_diff_lb <- exp(b1_lb)
ratio_diff_ub <- exp(b1_ub)
# Convert ratio to % change
pct_diff <- (ratio_diff - 1) * 100
pct_diff_lb <- (ratio_diff_lb - 1) * 100
pct_diff_ub <- (ratio_diff_ub - 1) * 100
# Put into a simple table
diff_table <- data.frame(
comparison = "Treated vs Non-treated",
pct_diff = pct_diff,
pct_lb = pct_diff_lb,
pct_ub = pct_diff_ub
)
diff_table# LOW
s <- summary(mP_lnr)
# Extract the moderator row (difference)
b1_est <- s$b[1] # lnRR difference: present - absent
b1_lb <- s$ci.lb[1] # lower CI
b1_ub <- s$ci.ub[1] # upper CI
# Convert lnRR difference to ratio
ratio_diff <- exp(b1_est)
ratio_diff_lb <- exp(b1_lb)
ratio_diff_ub <- exp(b1_ub)
# Convert ratio to % change
pct_diff <- (ratio_diff - 1) * 100
pct_diff_lb <- (ratio_diff_lb - 1) * 100
pct_diff_ub <- (ratio_diff_ub - 1) * 100
# Put into a simple table
diff_table <- data.frame(
comparison = "Treated vs Non-treated",
pct_diff = pct_diff,
pct_lb = pct_diff_lb,
pct_ub = pct_diff_ub
)
diff_tablemP_lnr2 <- rma(yi, vi, mods = ~ season, data = es_lnr, method = "REML")
mP_lnr2
Mixed-Effects Model (k = 19; tau^2 estimator: REML)
tau^2 (estimated amount of residual heterogeneity): 0.0062 (SE = 0.0023)
tau (square root of estimated tau^2 value): 0.0789
I^2 (residual heterogeneity / unaccounted variability): 95.28%
H^2 (unaccounted variability / sampling variability): 21.21
R^2 (amount of heterogeneity accounted for): 3.27%
Test for Residual Heterogeneity:
QE(df = 17) = 226.4658, p-val < .0001
Test of Moderators (coefficient 2):
QM(df = 1) = 1.6070, p-val = 0.2049
Model Results:
estimate se zval pval ci.lb ci.ub
intrcpt 0.0332 0.0271 1.2231 0.2213 -0.0200 0.0864
seasonsecond 0.0475 0.0375 1.2677 0.2049 -0.0260 0.1210
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1s2 <- summary(mP_lnr2)
# Extract the moderator row (difference)
b1_est <- s2$b[2] # lnRR difference: present - absent
b1_lb <- s2$ci.lb[2] # lower CI
b1_ub <- s2$ci.ub[2] # upper CI
# Convert lnRR difference to ratio
ratio_diff <- exp(b1_est)
ratio_diff_lb <- exp(b1_lb)
ratio_diff_ub <- exp(b1_ub)
# Convert ratio to % change
pct_diff <- (ratio_diff - 1) * 100
pct_diff_lb <- (ratio_diff_lb - 1) * 100
pct_diff_ub <- (ratio_diff_ub - 1) * 100
# Put into a simple table
diff_table2 <- data.frame(
comparison = "First vs Second",
pct_diff = pct_diff,
pct_lb = pct_diff_lb,
pct_ub = pct_diff_ub
)
diff_table2if (max(pred_tab$gain_hat) < 20) {
pred_tab <- pred_tab %>%
mutate(across(c(gain_hat, ci_lb, ci_ub, se), ~ .x * 1000))
}Pressure
Gcrit <- 1000 * cost / price
econ_tab <- pred_tab %>%
mutate(
Net = gain_hat * price / 1000 - cost,
Net_lb = ci_lb * price / 1000 - cost,
Net_ub = ci_ub * price / 1000 - cost,
Prob_Pos = 1 - pnorm((Gcrit - gain_hat) / se)
)
econ_tabsack_kg <- 60
econ_tab2 <- econ_tab %>%
mutate(
Prob_5s = 1 - pnorm((5 * sack_kg - gain_hat) / se),
Prob_10s = 1 - pnorm((10 * sack_kg - gain_hat) / se),
Prob_15s = 1 - pnorm((15 * sack_kg - gain_hat) / se)
)
econ_tab2econ_summary <- econ_tab2 %>%
transmute(
pressure,
net_return_R_ha = round(Net, 1),
prob_net_positive = round(Prob_Pos, 3)*100,
prob_5_sacks = round(Prob_5s, 3)*100,
prob_10_sacks = round(Prob_10s, 3)*100,
prob_15_sacks = round(Prob_15s, 3)*100
)
econ_summarySeason
econ_tab2 <- pred_tab2 %>%
mutate(
Net = gain_hat * price / 1000 - cost,
Net_lb = ci_lb * price / 1000 - cost,
Net_ub = ci_ub * price / 1000 - cost,
Prob_Pos = 1 - pnorm((Gcrit - gain_hat) / se)
)
econ_tab2sack_kg <- 60
econ_tab3 <- econ_tab2 %>%
mutate(
Prob_5s = 1 - pnorm((5 * sack_kg - gain_hat) / se),
Prob_10s = 1 - pnorm((10 * sack_kg - gain_hat) / se),
Prob_15s = 1 - pnorm((15 * sack_kg - gain_hat) / se)
)
econ_tab3econ_summary2 <- econ_tab3 %>%
transmute(
season,
net_return_R_ha = round(Net, 1),
prob_net_positive = round(Prob_Pos, 3)*100,
prob_5_sacks = round(Prob_5s, 3)*100,
prob_10_sacks = round(Prob_10s, 3)*100,
prob_15_sacks = round(Prob_15s, 3)*100
)
econ_summary2trial_disease <- hyb_app %>%
filter(application == "check") %>%
group_by(trial, season) %>%
summarise(across(all_of(diseases), mean, na.rm = TRUE), .groups = "drop") %>%
mutate(sev_sum = rowSums(across(all_of(diseases)), na.rm = TRUE)) %>%
arrange(sev_sum)
trial_diseaseheat_dat <- trial_disease %>%
mutate(trial_ord = factor(trial, levels = trial)) %>%
pivot_longer(all_of(diseases), names_to = "disease", values_to = "severity") %>%
bind_rows(
trial_disease %>%
mutate(trial_ord = factor(trial, levels = trial),
disease = "Total",
severity = sev_sum)
) %>%
mutate(disease = factor(disease, levels = c(diseases, "Total")))
heat_datheat_dat %>%
filter(!is.nan(severity)) %>%
group_by(disease) %>%
summarise(
mean = mean(severity))library(dplyr)
x_map <- c(
comum = "CR",
branca = "TR",
diplo = "DLS",
turc = "NCLB",
bipol = "SCLF",
antrac = "AL",
poli = "SR",
cercos = "GLS"
)
y_map <- c(
"2019_Pato Branco_first" = "2019_PB_1",
"2018_Campo Mourão_second" = "2018_CM_2",
"2019_Londrina_second" = "2019_L_2",
"2016_Pato Branco_first" = "2016_PB_1",
"2018_Pato Branco_first" = "2018_PB_1",
"2016_Campo Mourão_first" = "2016_CM_1",
"2020_Pato Branco_first" = "2020_PB_1",
"2018_Londrina_second" = "2018_L_2",
"2016_Londrina_second" = "2016_L_2",
"2020_Londrina_second" = "2020_L_2",
"2020_Campo Mourão_first" = "2020_CM_1",
"2018_Campo Mourão_first" = "2018_CM_1",
"2020_Campo Mourão_second" = "2020_CM_2",
"2017_Londrina_second" = "2017_L_2",
"2016_Campo Mourão_second" = "2016_CM_2",
"2017_Campo Mourão_first" = "2017_CM_1",
"2019_Campo Mourão_second" = "2019_CM_2",
"2019_Campo Mourão_first" = "2019_CM_1",
"2017_Campo Mourão_second" = "2017_CM_2"
)
heat_dat2 <- heat_dat %>%
mutate(
disease = recode(disease, !!!x_map),
trial_ord = recode(trial_ord, !!!y_map)
)
p_heat <- ggplot(heat_dat2, aes(x = disease, y = trial_ord, fill = severity,
label = round(severity, 2))) +
geom_tile(color = "white", linewidth = 0.2) +
geom_text(size = 5, color = "white") + # <- set text color here
labs(
x = "Disease in the non-treated",
y = "",
fill = "Severity (%)"
) +
scale_fill_viridis_c(option = "D", begin = 0.3, end = 1, na.value = "grey80") +
ggthemes::theme_few() +
theme(text = element_text(size = 14, face = "bold"),
axis.title = element_text(size = 16, face = "bold"),
strip.text = element_text(size = 16, face = "bold"),
title = element_text(face = "bold"),
legend.position = "right")
p_heat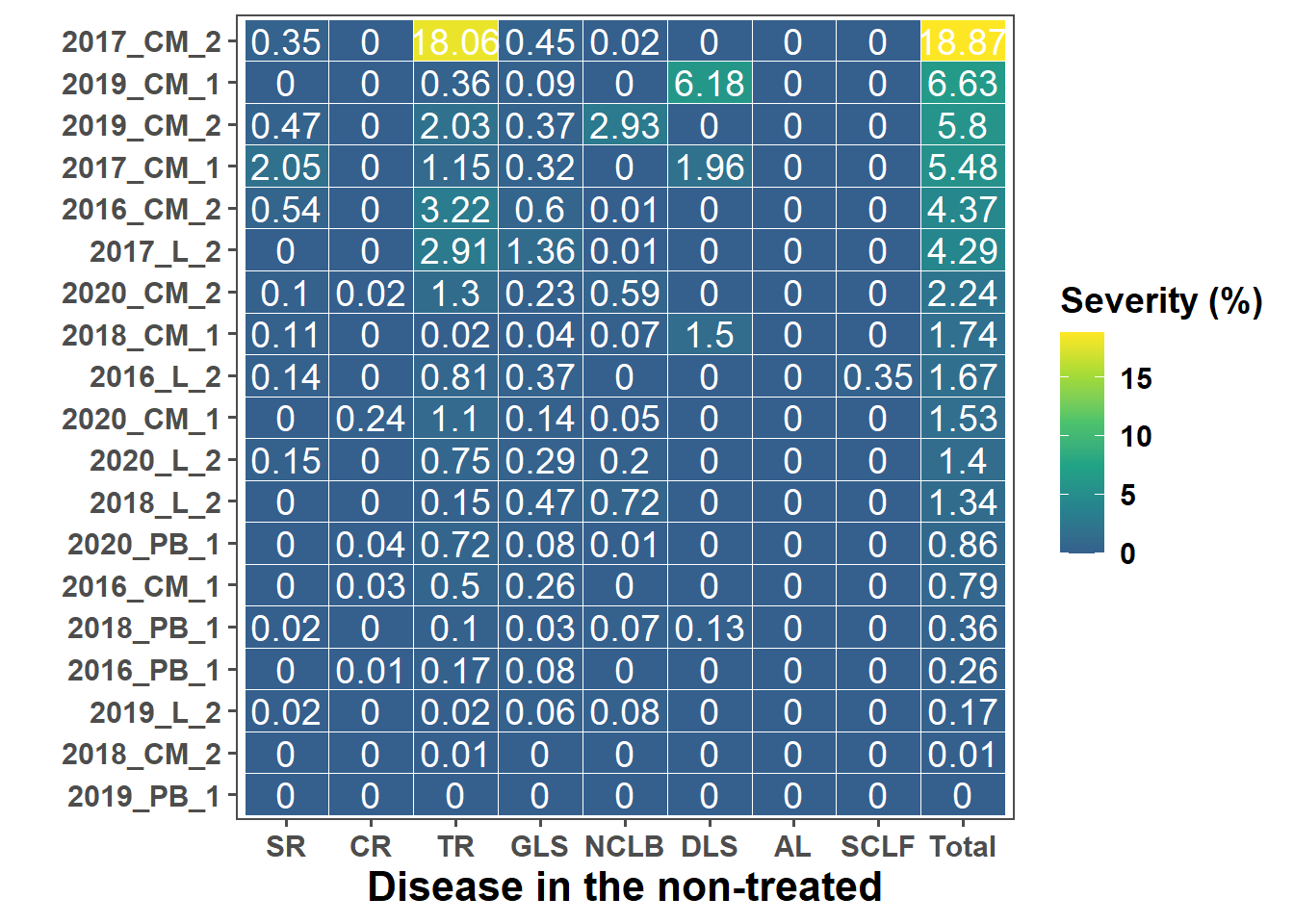
ggsave("fig/heat_disease.png", bg = "white", dpi = 600, height = 8, width = 12)p_net <- ggplot(econ_tab, aes(x = pressure, y = Net)) +
geom_hline(yintercept = 0, linewidth = 0.4) +
geom_col(width = 0.55, fill = "steelblue") +
geom_errorbar(aes(ymin = Net_lb, ymax = Net_ub), width = 0.1) +
labs(y = "Net return (R$ ha−1)", x = NULL) +
theme_half_open()
p_net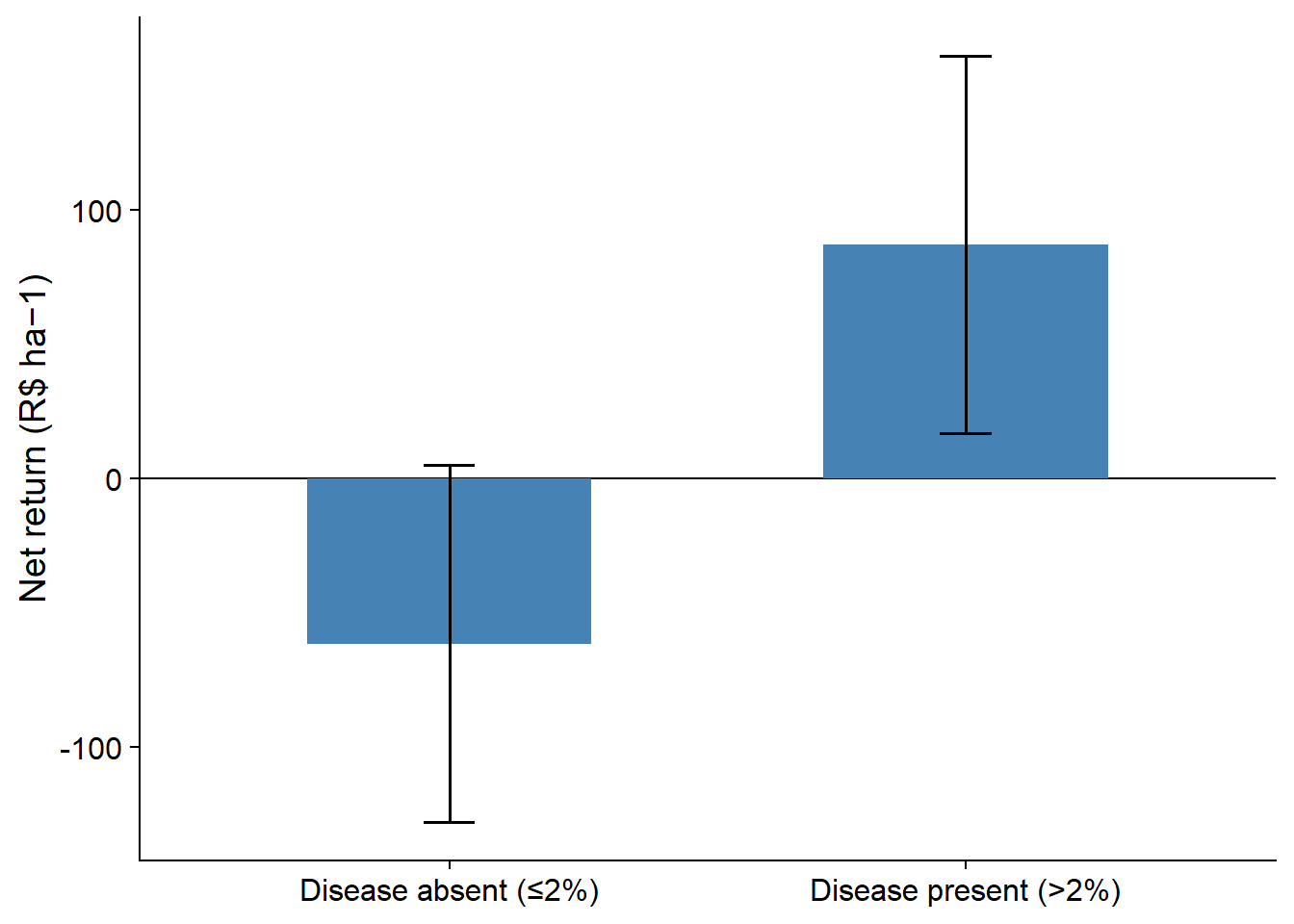
Yield
dados <- gsheet2tbl("https://docs.google.com/spreadsheets/d/1pUGN7bPHtENsO8eb-f8J1q6_pbMqm86bVnaNlgClleo/edit?usp=sharing")dados2 <- dados |>
mutate(trial = interaction(year, location, season, hybrid))
dados3 <- dados2 |>
dplyr::group_by(trial, year, location, season, hybrid, technology, application) |>
dplyr::summarize(sev = mean(severity),
yld = mean(yield2),
sd_sev = sd(severity),
sd_yld = sd(yield2),
n = n()
)
dados_meta <- dados3 %>%
dplyr::select(trial, year, location, season, hybrid, technology, application, yld, sd_yld, n, sev) %>%
pivot_wider(
names_from = application,
values_from = c(yld, sd_yld,n, sev),
names_sep = "_"
) %>%
dplyr::mutate(
study_id = paste(trial, year, location, season, hybrid, sep = "_"),
pressao = ifelse(sev_check > 10, "high", "low"),
yi = yld_treated - yld_check,
sei = sqrt((sd_yld_treated^2 / n_treated) + (sd_yld_check^2 / n_check))
) %>%
drop_na(yi, sei)
dados_metalibrary(ggthemes)
metaa = dados_meta %>%
filter(sev_check >=5) %>%
filter(yld_check <= 12000)
median(metaa$sev_check)[1] 10.28333yield_plot = metaa %>%
ggplot(aes(yld_check))+
geom_histogram(color = "white",fill = "black", bins = 10)+
scale_x_continuous(breaks = c(5000,7500,10000,12500),
limits = c(5000, 12500))+
facet_wrap(~season,labeller = as_labeller(c(
"first" = "",
"second" = " "
)))+
theme_few()+
labs(x = "Yield (kg/ha)",
y = "Frequency")+
theme(text = element_text(size = 8, face = "bold"),
axis.text.x = element_text(size = 8, face = "bold", angle = 45, vjust = 0.5),
axis.text.y = element_text(size = 8, face = "bold"),
axis.title = element_text(size = 12, face = "bold"),
strip.text = element_text(size = 12))
yield_plot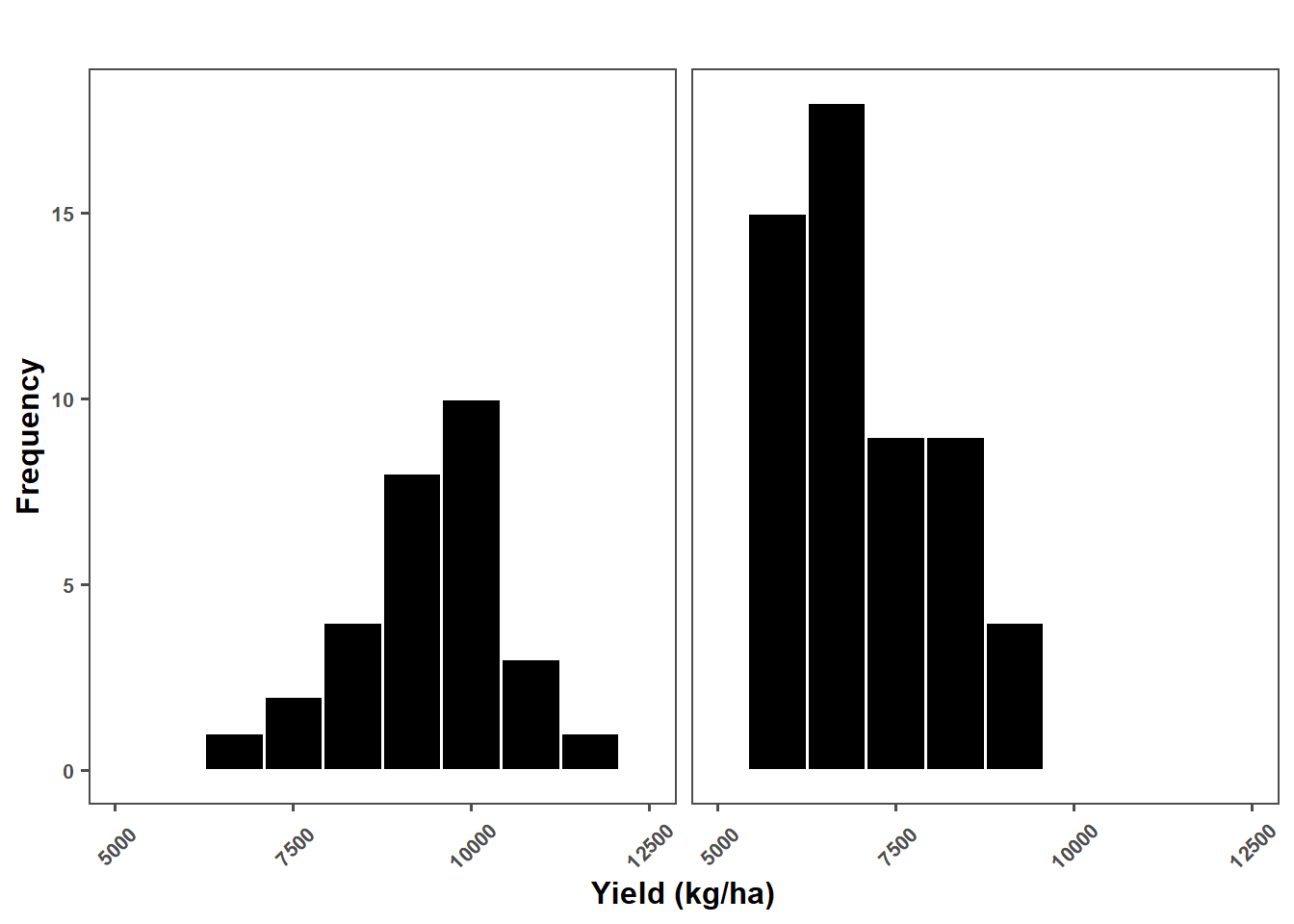
Severity
sev_plot = metaa %>%
ggplot(aes(sev_check))+
geom_histogram(color = "white",fill = "black", bins = 10)+
facet_wrap(~season,labeller = as_labeller(c(
"first" = "First",
"second" = "Second"
)))+
theme_few()+
labs(x = "Severity (%)",
y = "Frequency")+
theme(text = element_text(size = 8, face = "bold"),
axis.text.x = element_text(size = 8, face = "bold", angle = 45, vjust = 0.5),
axis.text.y = element_text(size = 8, face = "bold"),
axis.title = element_text(size = 12, face = "bold"),
strip.text = element_text(size = 12))
sev_plot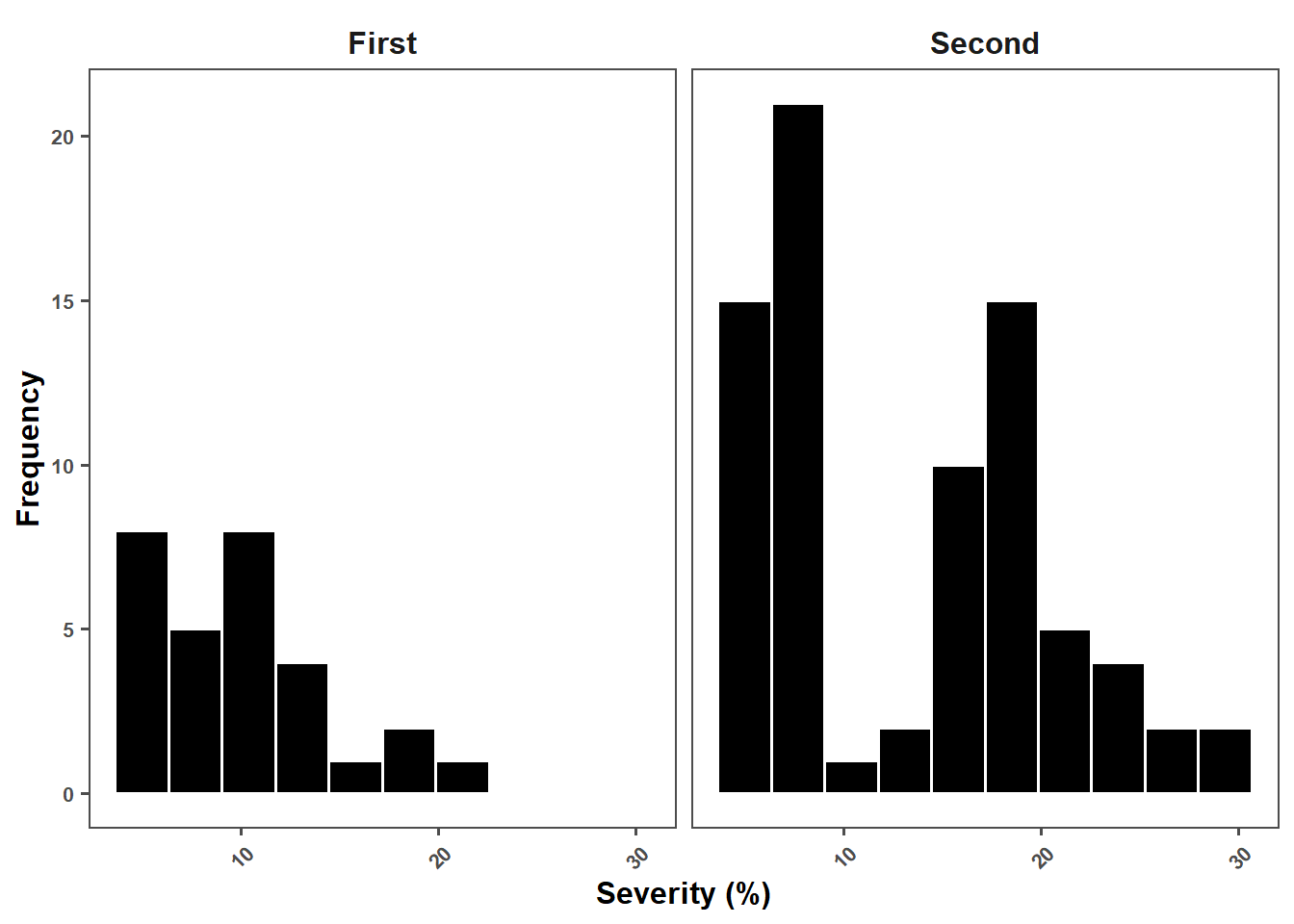
Map
library(dplyr)
library(ggplot2)
library(ggrepel)
library(maps)
# Número de híbridos por location + coordenadas
loc_hybrids <- dados %>%
group_by(location, season, lat, lon) %>%
summarise(n_hybrids = n_distinct(hybrid), .groups = "drop")
loc_hybrids$lat = NULL
loc_hybrids$lon = NULL
loc_hybrids$lat = c(-24.09,-24.09,-23.356,-26.1255)
loc_hybrids$lon = c(-52.356389,-52.356389,-51.158889,-52.65)
library(scales)
library(ggspatial)
library(gsheet)
library(ggrepel)
library(cowplot)
library(rnaturalearth)
library(rnaturalearthdata)
library(sf)
library(tidyverse)
#install.packages("devtools") # I guess you also need this
#devtools::install_github("ropensci/rnaturalearthhires")
BRA = ne_states(
country = "Brazil",
returnclass = "sf"
)
library(geobr)
municipios <- read_municipality(code_muni = "all", year = 2020)
municipios = municipios %>% filter(name_state == c("Paraná"))
states <- filter(BRA,
name_pt == "Paraná")
unique(BRA$name_pt) [1] "Rio Grande do Sul" "Roraima" "Pará"
[4] "Acre" "Amapá" "Mato Grosso do Sul"
[7] "Paraná" "Santa Catarina" "Amazonas"
[10] "Rondônia" "Mato Grosso" "Maranhão"
[13] "Piauí" "Ceará" "Rio Grande do Norte"
[16] "Paraíba" "Pernambuco" "Alagoas"
[19] "Sergipe" "Bahia" "Espírito Santo"
[22] "Rio de Janeiro" "São Paulo" "Goiás"
[25] "Federal" "Minas Gerais" "Tocantins" states = states %>%
mutate(id = case_when(
name_pt == "Paraná" ~ "PR"))
SUL = ne_states(
country = c("Argentina", "Uruguay", "Paraguay", "Colombia", "Bolivia"),
returnclass = "sf")
br_sf <- ne_states(geounit = "brazil",
returnclass = "sf")
library(ggplot2)
library(cowplot)
library(ggplot2)
library(ggrepel)
library(dplyr)
library(ggthemes)
library(ggspatial)
loc_hybrids <- loc_hybrids[-1, ]
map3 <- loc_hybrids |>
ggplot() +
# Mapa base do Brasil
geom_sf(data = BRA, fill = "white", color = "gray60", size = 0.4) +
# Destacar o Paraná
geom_sf(data = subset(BRA, postal %in% c("PR")),
fill = NA, color = "black", size = 1) +
# Siglas dos estados
geom_text(data = states, aes(x = longitude, y = latitude, label = id),
size = 5, hjust = 0.8, color = "black", fontface = "bold") +
# Pontos das localidades
geom_point(data = loc_hybrids, aes(x = lon, y = lat),
alpha = 0.8) +
# Nomes das cidades com linhas
geom_text_repel(
data = loc_hybrids,
aes(x = lon, y = lat, label = location),
size = 4,
fontface = "bold",
color = "black",
min.segment.length = 0,
box.padding = 0.3,
segment.color = "gray30",
segment.size = 0.4
) +
# Tema e ajustes de mapa
theme_minimal_grid() +
coord_sf(xlim = c(-58, -45), ylim = c(-27, -22), expand = FALSE) +
# Legenda personalizada
guides(size = FALSE,
color = guide_colorbar(title = "Severity (%)",
direction = "horizontal",
title.position = "top")) +
# Estilo da legenda e borda do painel
theme(
legend.position = c(0.9, 0.01),
legend.justification = c(1, 0),
legend.direction = "horizontal",
legend.title.align = 0.5,
panel.border = element_rect(color = "black", fill = NA, size = 1),
axis.title = element_text(face = "bold")
) +
# Adiciona seta de norte
annotation_north_arrow(location = "br", which_north = "true",
pad_x = unit(5.86, "in"), pad_y = unit(0.3, "in"),
style = north_arrow_fancy_orienteering,
height = unit(1.2, "cm"), width = unit(1.1, "cm")) +
# Barra de escala
annotation_scale(location = "br",
width_hint = 0.1,
height = unit(0.2, "cm"),
text_cex = 0.6,
pad_x = unit(5.8, "in"),
pad_y = unit(0.2, "in")) +
# Eixos
labs(x = "Longitude", y = "Latitude")
map3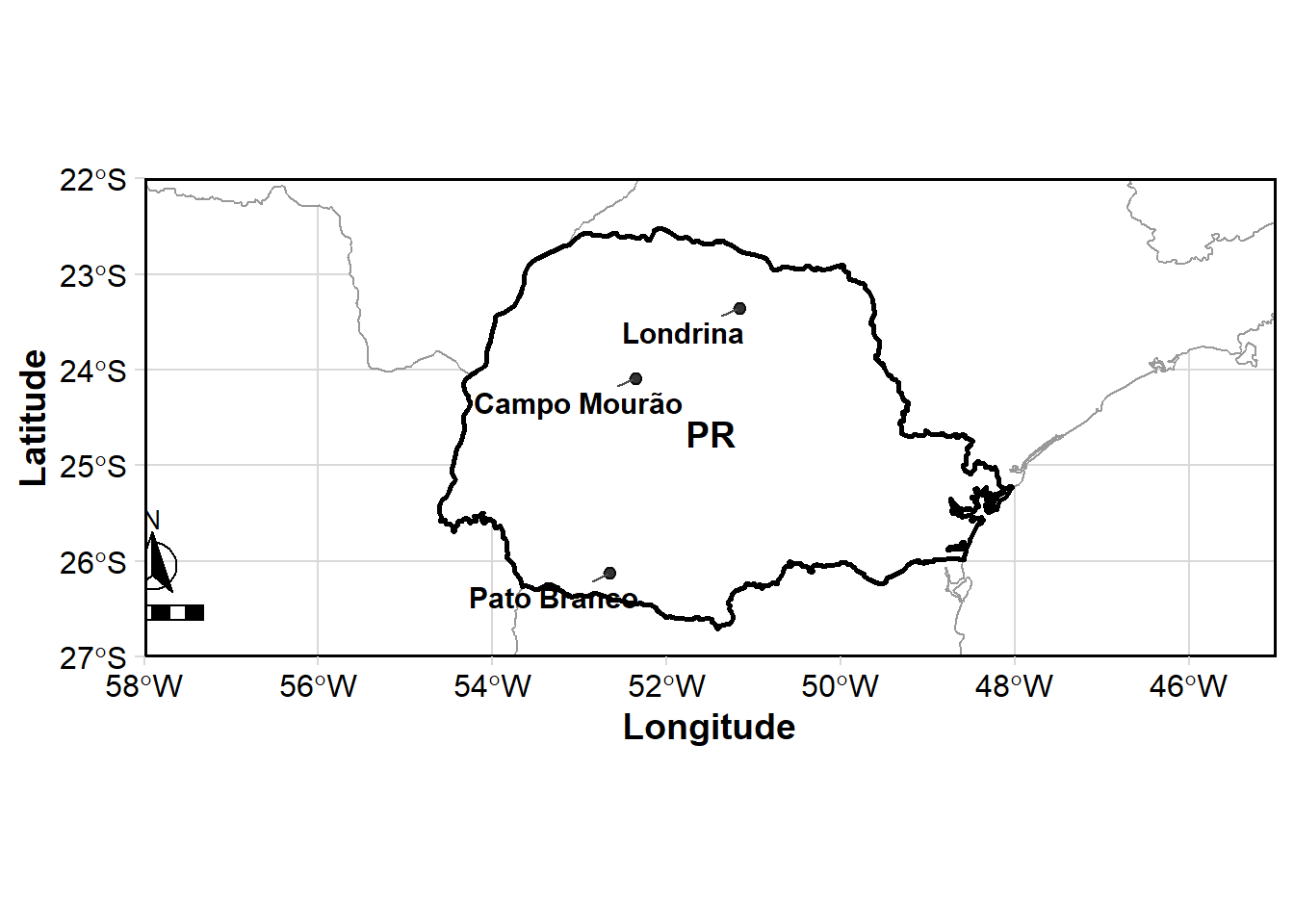
pr_municipios <- read_municipality(code_muni = "all", year = 2020) %>%
filter(name_state == "Paraná")
BRA <- ne_states(country = "Brazil", returnclass = "sf")
pr_estado <- BRA %>% filter(name_pt == "Paraná")
map2 = loc_hybrids #|> filter(estado == "PR")
mapa_brasil <- ggplot() +
geom_sf(data = BRA, fill = "white", color = "black", size = 0.2) +
geom_sf(data = subset(BRA, postal %in% c("PR")),
fill = "gray40", color = "black", size = 0.6) +
theme_map() +
theme(
panel.background = element_rect(fill = "white", color = NA),
plot.background = element_rect(fill = "white", color = NA),
panel.border = element_rect(color = "black", fill = NA, size = 1)) library(patchwork)
mapp = map3 +
inset_element(mapa_brasil,
left = 0.8, bottom = 0.5,
right = 0.99, top = 0.95) library(cowplot)
right_panel <- plot_grid(
sev_plot, yield_plot,
ncol = 1,
labels = c("(b)", "(c)"),
label_size = 14,
label_fontface = "bold",
align = "v"
)
final_plot <- plot_grid(
mapp, right_panel,
ncol = 2,
rel_widths = c(2, 1),
labels = c("(a)",""),
label_size = 14,
label_fontface = "bold",
label_x = 0.02,
label_y = 0.98,
align = "h"
)
final_plot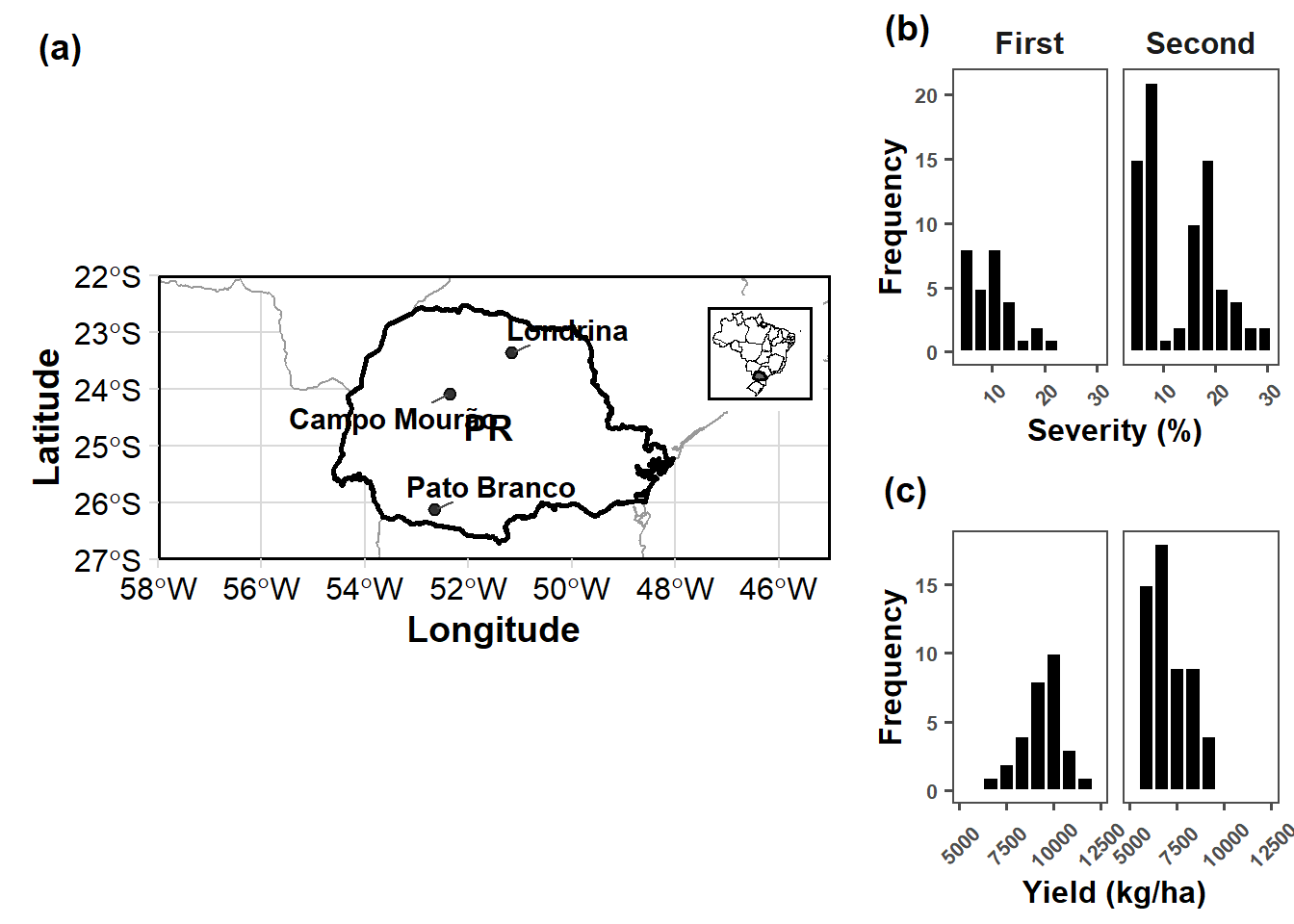
ggsave("fig/map.png", height=4, width=10, dpi = 600, bg = "white")